dataset - Is there a way to convert smiles format to TUdataset
4.6 (330) · $ 12.50 · In stock
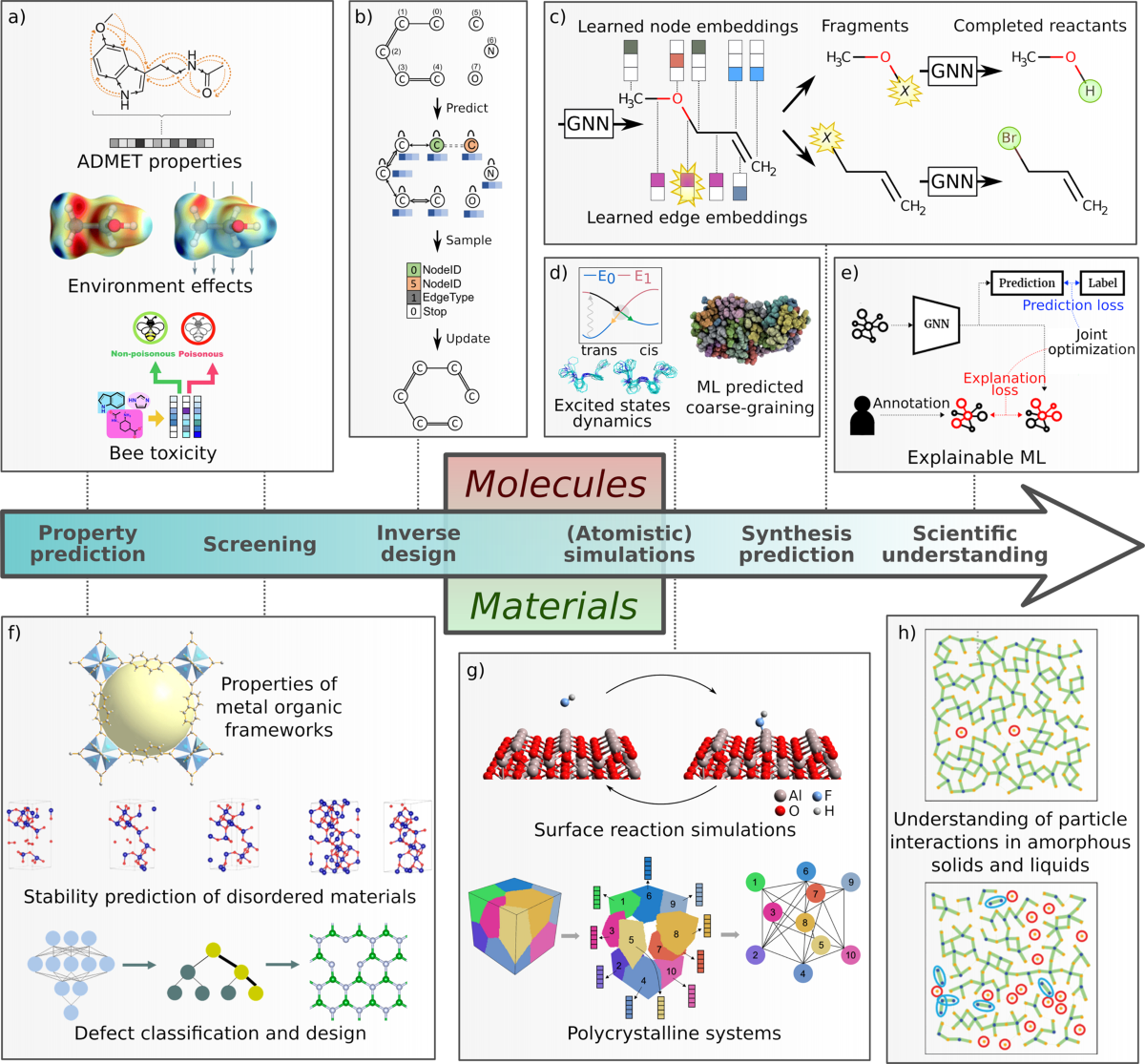
Graph neural networks for materials science and chemistry
GitHub - RohanV01/Molecule_format_converter: This jupyter notebook let's you convert sdf to pdb or SMILES to pdbqt file formats for a batch of compounds to perform cheminformatics and drug discovery projects
sdftosmi.py: Convert multiple ligands/compounds in SDF format to SMILES. — Bioinformatics Review

PDF] Multi-caption Text-to-Face Synthesis: Dataset and Algorithm

PDF) Visual Analytics For Machine Learning: A Data Perspective Survey
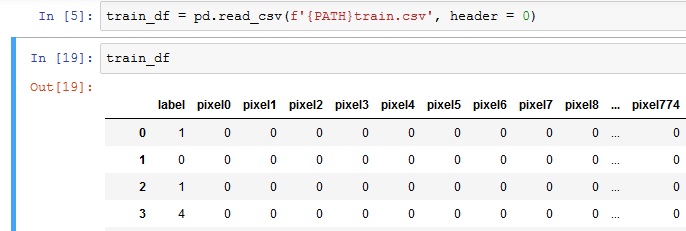
How to use Kaggle's MNIST data with ImageClassifierData? - Beginner (2018) - fast.ai Course Forums

PDF) EPIC: Graph Augmentation with Edit Path Interpolation via Learnable Cost

How to Convert Data Formats for Visualization

Converting dataset to json HELP - General Discussion - Inductive Automation Forum

sdftosmi.py: Convert multiple ligands/compounds in SDF format to SMILES. — Bioinformatics Review

PDF) EPIC: Graph Augmentation with Edit Path Interpolation via Learnable Cost

How to convert bundle SMILES to .sdf file format #Molecular_Docking #ChemistryStudio